September 2022 Clinical Case of the Month
Title: Selection of haploidentical stem cell donors
Submitted by: Syed Ali Abutalib and Nicolaus Kröger
Physicians expert perspective: Katharina Fleischhauer
A 46-year-old female is scheduled to undergo T-cell replete HLA-haploidentical hematopoietic cell transplantation (haplo-HCT) using post-transplant cyclophosphamide (PTCy), tacrolimus, and mycophenolate mofetil for acute myeloid leukemia in second remission.
Two potential HLA-haploidentical family donors (haplo-donors) are available; both are sons matched for ABO blood group and cytomegalovirus (CMV) serology. The recipient does not have donor-specific antibodies (DSA) against the mismatched HLA in either of the two donors.
- Donor 1: HLA-DQB1 is matched and the 5 other loci (HLA-A, B, C, DRB1, DPB1) are mismatched on the unshared haplotype with the patient. There is a B-leader match and a DPB1 T-cell epitope (TCE) nonpermissive mismatch.
- Donor 2: HLA-DRB1 and HLA-A are matched and the 4 other loci (HLA-B, C, DQB1, DPB1) are mismatched on the unshared haplotype with the patient. There is a B-leader mismatch and a DPB1 TCE permissive mismatch.
Which of the following statements is correct?
A. Donor 1 is to be preferred over Donor 2
B. Donor 2 is to be preferred over Donor 1
C. Both Donors are equivalent and can be selected based on logistic considerations
Expert Perspective by Katharina Fleischhauer
Haplo-donors share by genetic inheritance one of their two HLA-A, -B, -C, -DRB1, -DQB1, -DPB1 haplotypes with the patient, while the other haplotype is unshared. Therefore, they are commonly mismatched with the patient for 6 out of 12 HLA alleles, although this number can vary (Figure). Genetic recombination, occurring with a frequency of up to 1% if patient and donor are not related by direct descent1, can lead to the presence of mismatches also on the shared haplotype. Both candidate haplo-donors for our patient are sons, obviating the necessity to exclude recombination, which is however recommended for all haplo-donors other than mother, father, son or daughter.
Incidental sharing of alleles on the unshared haplotype, or homozygosity for individual loci, lead to less than 6 HLA mismatches in haploidentical pairs (Figure). Donor 1 has 5 mismatches with our patient, while donor 2 has 4. However, in haplo-HCT under graft-versus-host disease (GVHD) prophylaxis by PTCy, the number of HLA mismatches on the unshared haplotype was not predictive of GVHD or survival2. Rather, defined locus-specific characteristics of HLA mismatches informed survival (Figure), which was improved by B-leader matches3, DRB1 mismatches in graft-versus-host (GVH) direction, and nonpermissive DPB1 T-cell epitope (TCE) mismatches4. For our patient, all three favorable characteristics are present in donor 1 but not in donor 2, resulting in 3-year survival probabilities of 0.78 and 0.57, respectively, for this patient (http://haplodonorselector.b12x.org/v1.0/). Biologically, the favorable impact of B-leader matches and HLA class II mismatches might be explained by NK-5 and T-cell6 mediated alloreactivity, respectively, in the PTCy setting which appears to dampen the immunogenicity of HLA alloantigens7,8.
HLA mismatches can also be recognized by DSA, which can interfere with engraftment9. Therefore, all patients scheduled for haplo-HCT should be tested for DSA, and haplo-donors carrying DSA targets should be avoided. In our patient, no DSA were detected against either of the two candidate donors.
In summary, both donors are equivalent for non-HLA factors (ABO, CMV), absence of DSA, and presence of an HLA-C mismatch, which has been associated with increased chronic GVHD2. However, they differ for the number and locus-specific characteristics of HLA mismatches. Answer C is therefore incorrect. Despite the lower number of HLA mismatches for donor 2, better locus-specific characteristics of the HLA mismatches determine better survival probabilities for donor 1.
Correct Answer – A
References:
1. Martin M, Mann D, Carrington M. Recombination rates across the HLA complex: use of microsatellites as a rapid screen for recombinant chromosomes. Hum Mol Genet 1995; 4(3): 423-8.
2. Fuchs EJ, McCurdy SR, Solomon SR, et al. HLA informs risk predictions after haploidentical stem cell transplantation with posttransplantation cyclophosphamide. Blood 2022; 139(10): 1452-68.
3. Petersdorf EW, Carrington M, O'hUigin C, et al. Role of HLA-B exon 1 in graft-versus-host disease after unrelated haemopoietic cell transplantation: a retrospective cohort study. Lancet Haematol 2020; 7(1): e50-60.
4. Fleischhauer K, Shaw BE, Gooley T, et al. Effect of T-cell-epitope matching at HLA-DPB1 in recipients of unrelated-donor haemopoietic-cell transplantation: a retrospective study. Lancet Oncol; 2012 13(4): 366-74.
5. Horowitz A, Djaoud Z, Nemat-Gorgani N, et al. Class I HLA haplotypes form two schools that educate NK cells in different ways. Sci Immunol 2016; 1(3): eaag1672.
6. Afzali B, Lechler RI, Hernandez-Fuentes MP. Allorecognition and the alloresponse: clinical implications. Tissue Antigens 2007; 69(6): 545-56.
7. Kanakry CG, Fuchs E, Luznik L. Modern approaches to HLA-haploidentical blood or marrow transplantation. Nat Rev Oncol 2016; 13(1): 10–24.
8. Fleischhauer K. Haplo-PtCy: adjusting the HLA barrier. Blood 2022; 139(10): 1431-3.
9. Ciurea S, Cao K, Fernadez-Vina M, et al. The European Society for Blood and Marrow Transplantation (EBMT) Consensus Guidelines for the Detection and Treatment of Donor Specific Anti-HLA Antibodies (DSA) in Haploidentical Hematopoietic Cell Transplantation. Bone Marrow Transplant 2018; 53(5): 521-34.
Figure: HLA mismatches in related haplo-HCT.
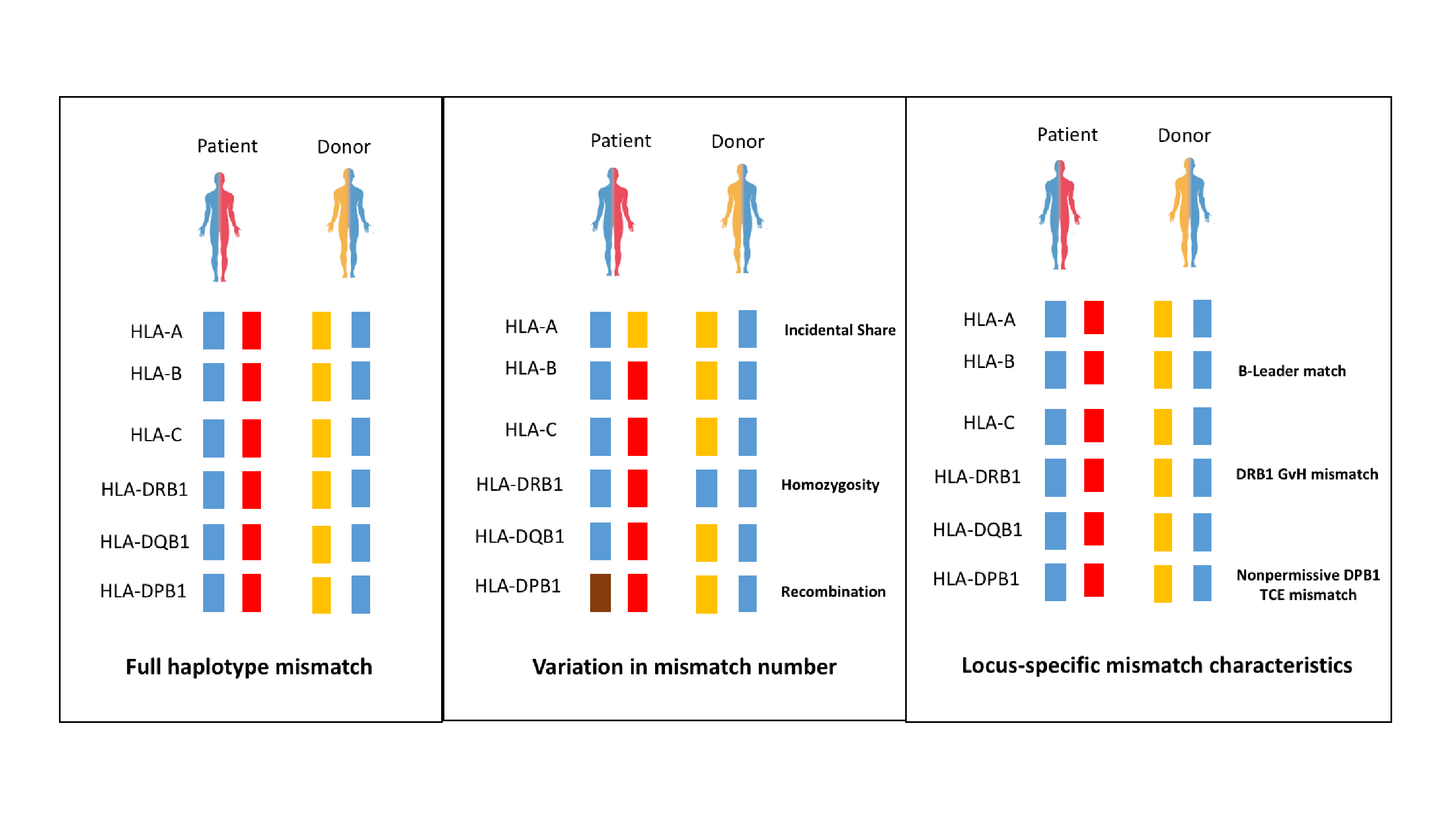
Figure legend: Left panel: patients and their haploidentical relatives share one full HLA haplotype by inheritance (blue), while the other haplotype is unshared (red and yellow), leading most commonly to 6 mismatched HLA alleles. Middle panel: Variation in the number of HLA mismatches can occur by incidental sharing of the allele of a given locus (here HLA-A) on the unshared haplotype between patient and donor, homozygosity at a given locus (here HLA-DRB1 of the donor), or, in donors not of direct descent, genetic recombination at a given locus (here HLA-DPB1). Note that in the latter case, all 4 alleles can be mismatched. Incidental sharing, homozygosity or recombination shown here in exemplary form for individual HLA loci might affect any of the 6 loci in the patient and/or the donor. Right panel: In haplo-HCT under PTCy, HLA mismatches with favorable locus-specific characteristics were identified2. These include match for the threonine/methionine dimorphism at position -21 of HLA-B (B-leader match)3, HLA-DRB1 mismatch in GVH direction (patient carrying a DRB1 allele not present in the donor, i.e. at least the patient heterozygous at DRB1, as in the examples in the right and middle panel), and non-permissive HLA-DPB1 TCE mismatch (TCE group disparity for DPB1)4. Predictions of survival probabilities based on these three favorable HLA mismatch characteristics and other relevant non-HLA parameters, can be obtained via the free online webtool at http://haplodonorselector.b12x.org/v1.0/, which however does not consider the presence of absence of DSA2.
Contacts
Syed Ali Abutalib, MD
Associate Director, Hematology and Cellular Therapy Program
Director, Clinical Apheresis Program
Cancer Treatment Centers of America, Zion, Illinois
Associate Professor, Rosalind Franklin University of Medicine and Science
Email: abutalib110@gmail.com
Nicolaus Kröger, MD
Professor and Medical Director of the Department of Stem Cell
Transplantation at the University Hospital Hamburg-Eppendorf, Germany
University Hospital Hamburg, Hamburg, Germany
Correspondence: Nicolaus Kröger, MD
Email: nkroeger@uke.uni-hamburg.de
Expert Perspective
Katharina Fleischhauer, M.D
Director of the Institute for Experimental Cellular Therapy
Professor at the Faculty of Medicine, University Duisburg-Essen, Germany
University Hospital Essen, Essen, Germany
Email: katharina.fleischhauer@uk-essen.de
Future Clinical Case of the Month
If you have a suggestion for future clinical case to feature, please contact Nicolaus Kröger.